Note
Go to the end to download the full example code.
Analysis of bivariate data#
The pymultifracs package can perform bivariate multifractal analysis,
and produces bivariate cumulants and structure functions. The purpose of
bivariate MFA is to determine the relationship between the regularity
fluctuations of a pair of time series.
We will be using pre-generated data to serve as an example of bivariate
multi-fractal time series. Let us first load the data; those example time
series are also found in the tests/data/ folder of the repository on
github.
Downloading data from 'https://github.com/neurospin/pymultifracs/raw/refs/heads/master/tests/data/DataSet_ssMF.mat' to file '/home/runner/.cache/bivar/c2970ce816c6ddbf5b2539426c580fc7-DataSet_ssMF.mat'.
SHA256 hash of downloaded file: 7027dfe19282c5802ff5e3bbab2fca7d9fb744c874a853d82883c2b0fed6fb57
Use this value as the 'known_hash' argument of 'pooch.retrieve' to ensure that the file hasn't changed if it is downloaded again in the future.
The data is a two-dimensional array, with both self-similarity and multifractal correlation.
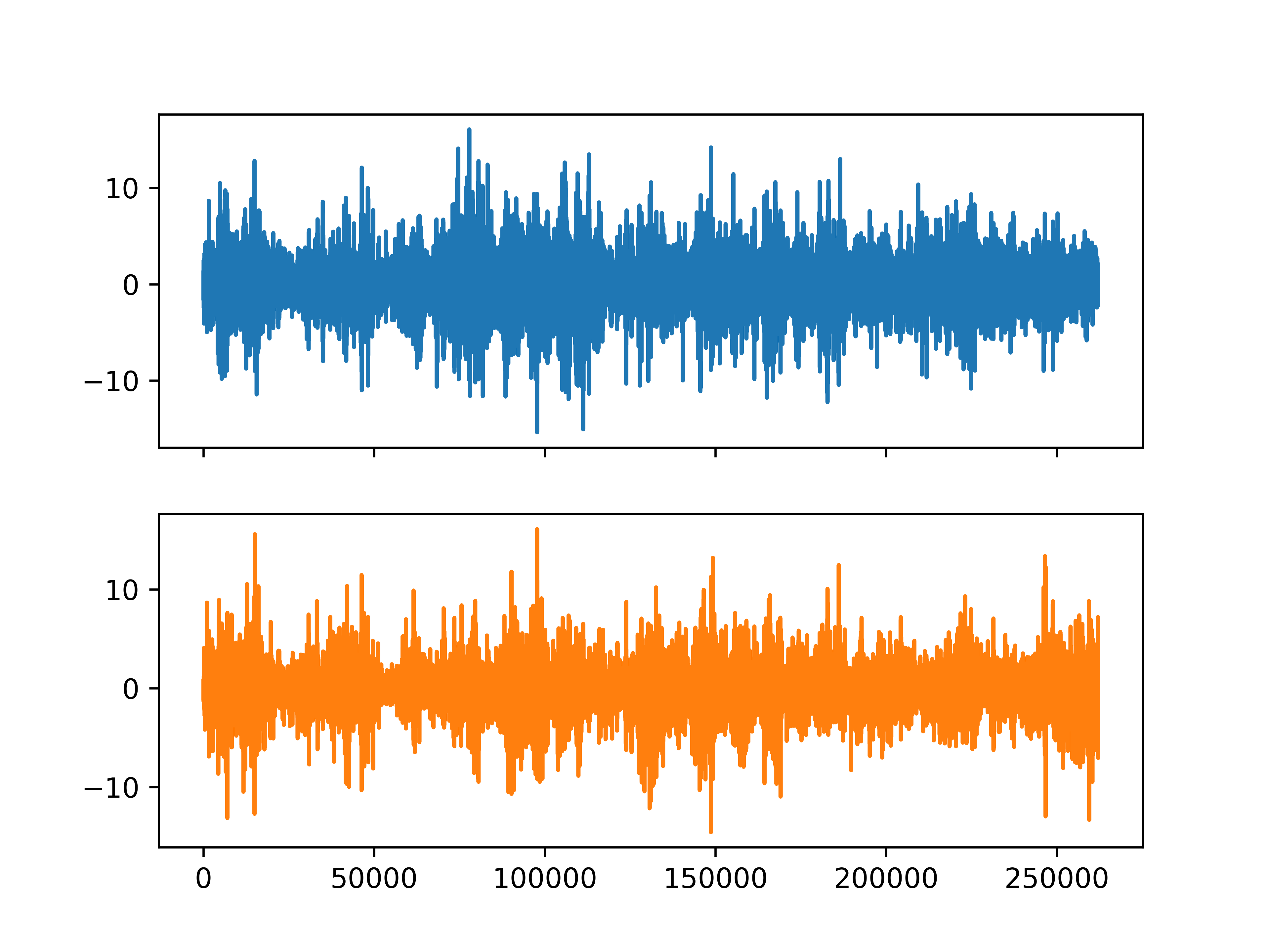
[<matplotlib.lines.Line2D object at 0x7f71df1a87d0>]
First we need wavelet_analysis() as ususal, to obtain p-leaders in
this case
from pymultifracs import wavelet_analysis
WTpL = wavelet_analysis(X).integrate(.75).get_leaders(2)
Then we use bivariate.bimfa() on the wavelet p-leaders. We provide
the same MRQ twice, in order to compute the estimates for all signal pairs.
# We can obtain the multifractal correlation matrix :math:`\rho_{mf}`:
print(pwt.cumulants.rho_mf.squeeze())
<xarray.DataArray (channel1: 2, channel2: 2)> Size: 32B
array([[1. , 0.71314563],
[0.71314563, 1. ]])
Coordinates:
m1 int64 8B 1
m2 int64 8B 1
Dimensions without coordinates: channel1, channel2
We can plot the structure function:
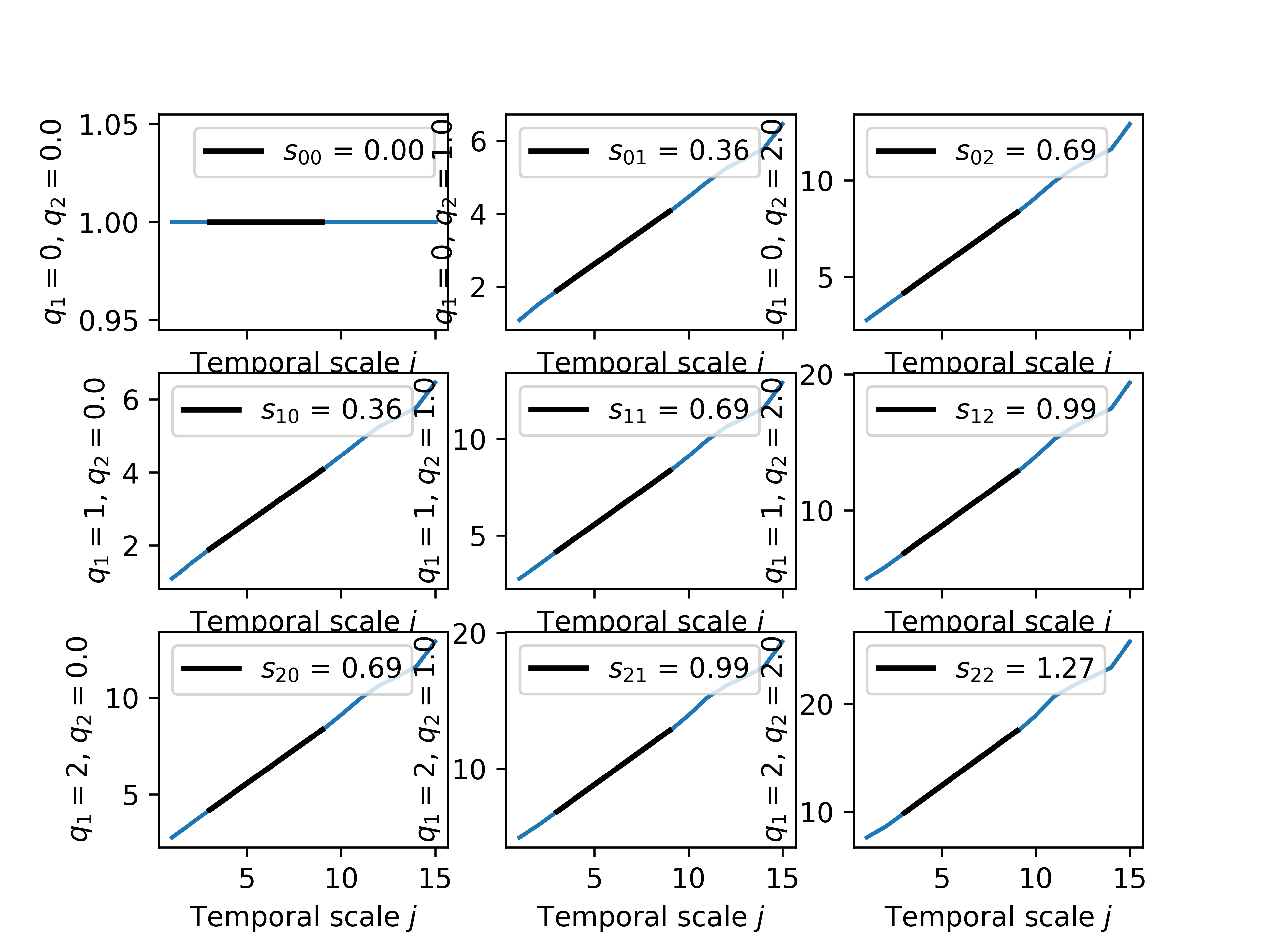
We can plot the bivariate cumulants:
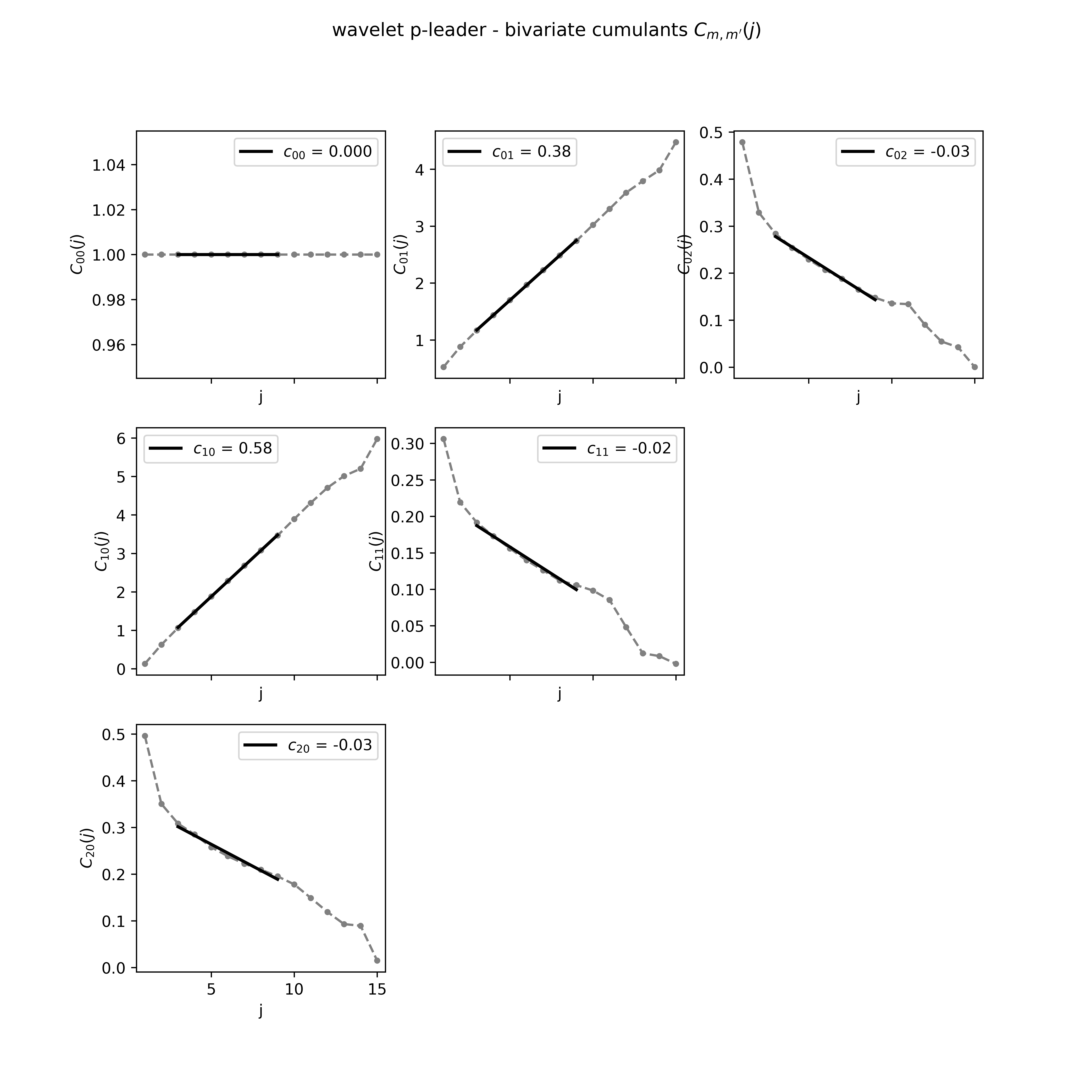
We can plot the Legendre spectrum reconstituted from the log-cumulants:
pwt.cumulants.plot_legendre(h_support=(.1, .95))
Total running time of the script: (0 minutes 8.841 seconds)